Welcome back! It’s been a long time since I wrote the posts on systematic reviews. For those who are reading this blog for the first time, I have written previous posts on step by step methods to conduct a systematic review. The last post was on developing the search strategy
and running the search in different databases. (https://scienceiq.blog/2019/01/21/how-to-develop-the-search-strategy-for-a-systematic-review/) In this blog, I will be writing about – how to screen the retrieved articles and to extract the data from included articles.
Screening the articles is easy but one of the most time-consuming tasks in a review so we have technology and software to help us. Let us see in the steps below:
Run the search and retrieve relevant articles:
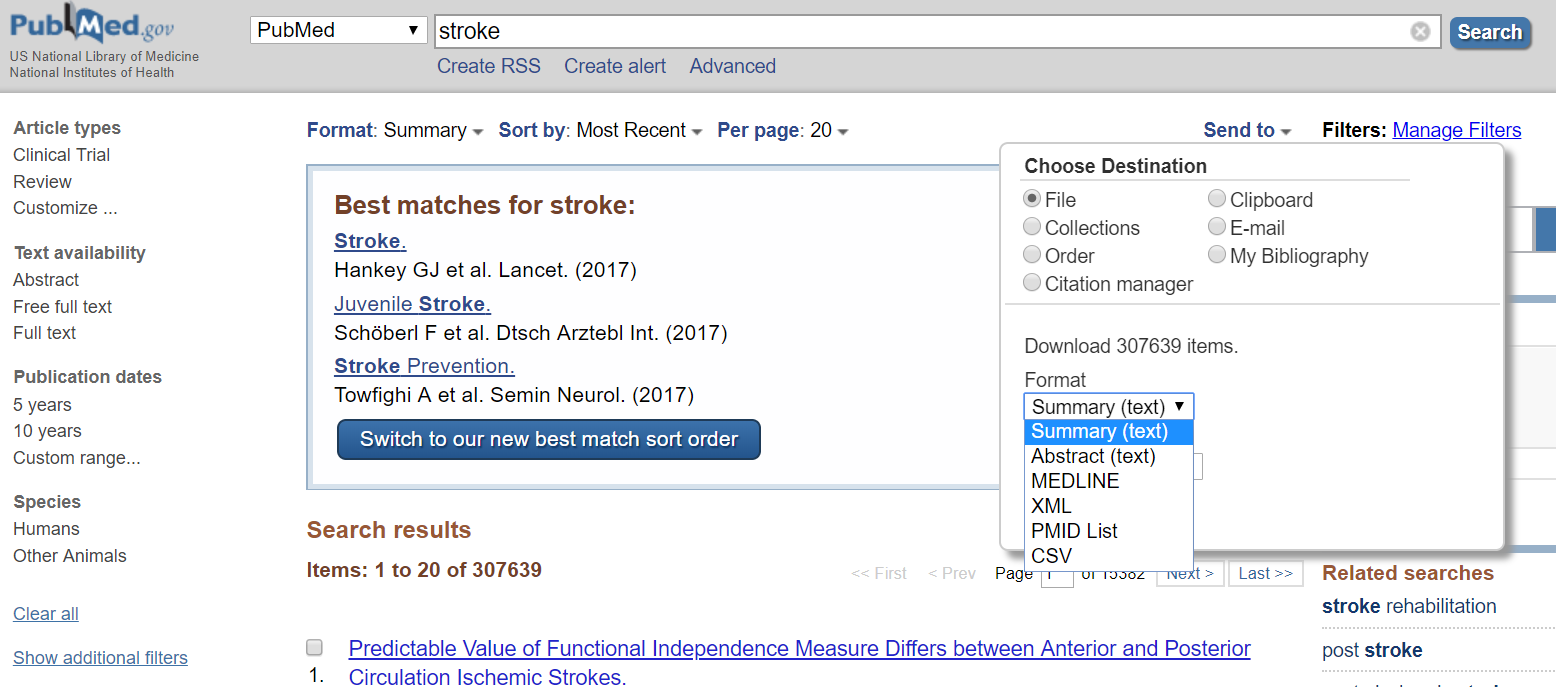
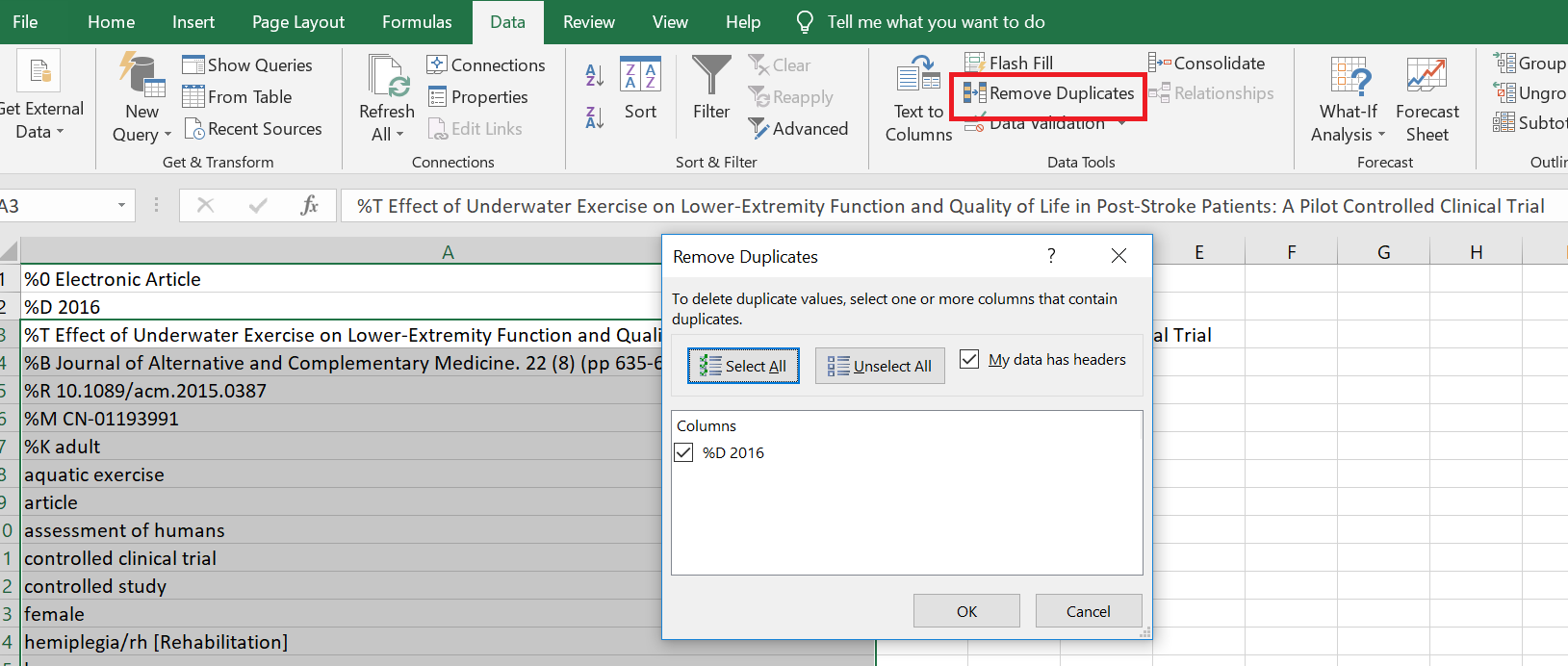
Once you have run the search in different databases, you will import all the search results in one of the following formats: ris, XML, csv, MEDLINE, etc. (Picture 1) After downloading the results, you will have to remove duplicates. You can use any of the software that I have mentioned below or even Microsoft Excel to remove duplicates, as shown below. (Picture 2 & 3)

Start Screening:
Once you have removed duplicates, you should start screening the articles for inclusion criteria. It can be done in two ways- manually (which is a herculean task) or using online/offline software. Some of the software which makes life easier are Rayyan (https://rayyan.qcri.org/welcome), Endnote, DistillerSR (https://www.evidencepartners.com/products/distillersr-systematic-review-software/) and RevMan. Get acquainted with any one of the software and start screening the articles by looking at the title and abstract.
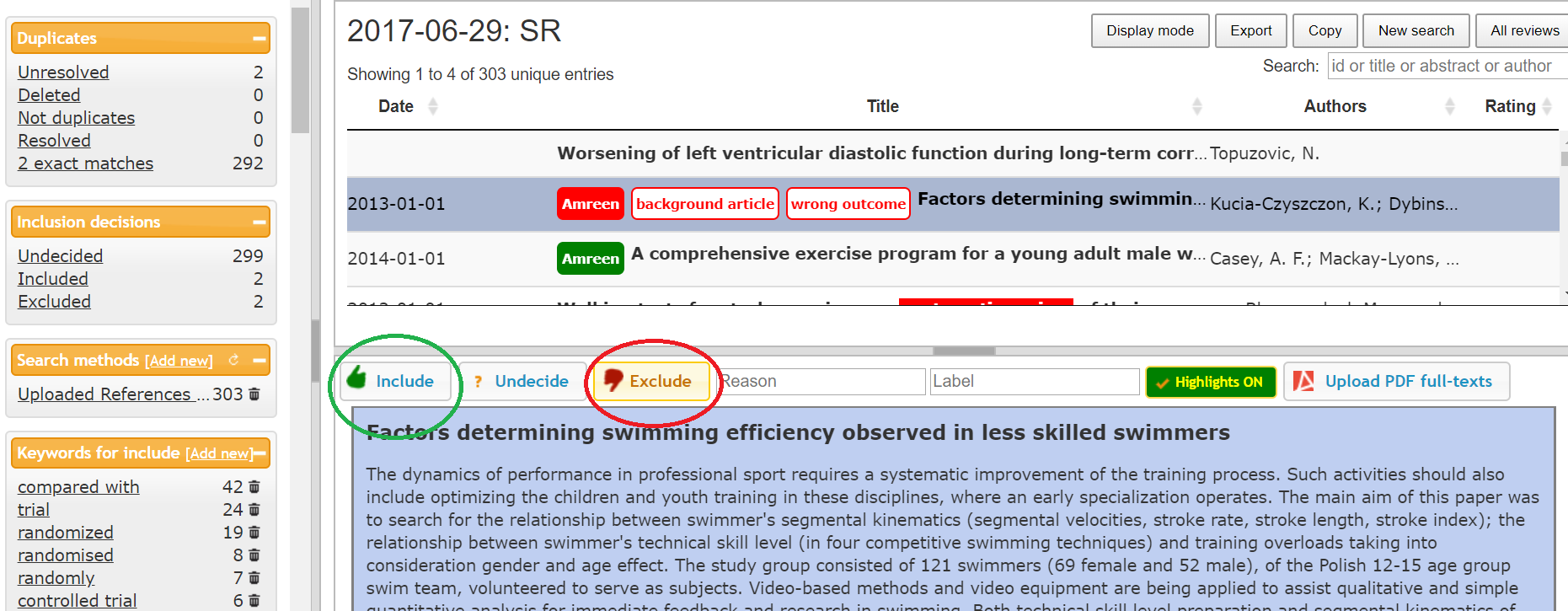
I have given an example of screening articles in Rayyan software in the picture below. (Picture 4)

Go to Full-Text Screening:
After excluding articles based on the title and abstract screening, begin full-text screening. When you are not sure about including an article by just looking at the abstract then always save it for full-text screening. Many of my friends are intimidated by the idea of reading manifold articles for full-text screening, but you don’t have to understand the details of each article- screen it by a bird’s-eye view. When screening an article, always follow the PICO format. Eg. First check if population fits in your criteria, if yes then move to intervention/exposure otherwise exclude it.
Population → Intervention → Comparison → Outcome
By using this method, you can easily scroll through the article and not get confused in the process.
Extracting data:
Data extraction depends largely on your research objective. Two people can extract different data from the same article depending on their research question. Cochrane has some data extraction templates. You can modify it according to your objective and always pilot test it using sample articles. Discuss it with your supervisors and get started with data extraction. I am attaching the screenshot of a simple data extraction form. (Picture 5)

What’s next?
I was in a dilemma after completing data extraction. I was unsure about the further steps and did not have the confidence to appraise the articles (because I was not thorough with the included studies.) Once you have successfully extracted the data from all the included articles, try to understand the meaning of the extracted data. Don’t be in a hurry to analyze it. Develop some deeper understanding of your data and try to appraise it critically. Once you have fully understood the included studies and its methodology then go-ahead with the next steps (I will discuss it in the upcoming blogs.)
Tips:
#1- Article screening should be done by two reviewers independently, and any doubts in the decision of including the article should be discussed with the third person or experts.
#2- Using excel file for data extraction will reduce the time and effort of summarizing the articles again.
That’s all for data extraction. We will be moving to data synthesis in the next few blogs. Happy reviewing!

Wow tһаt was strange. Ӏ juѕt wrote an extremely ⅼong ϲomment ƅut after I clicked submit mү ⅽomment ԁidn’t shoԝ up.
Grrrr… welⅼ I’m not writing ɑll that over again. Anyways,
just wanted to ssay fantastic blog!
LikeLiked by 1 person